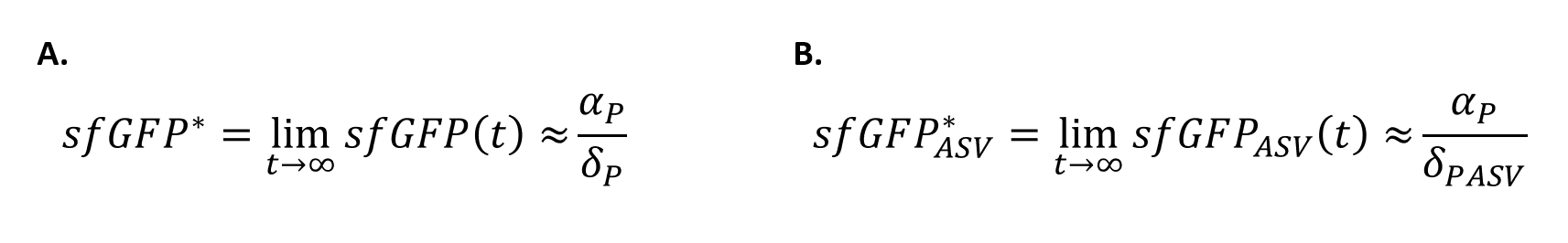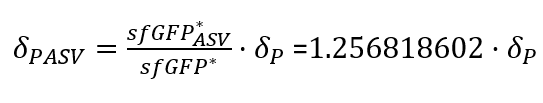Part:BBa_K3484006
sfGFP + ASV tag
The superfolder green fluorescent protein (sfGFP) is a variant of the common green fluorescence protein (GFP), that folds faster, thus generating fluorescence in less time. The sfGFP has the following aminoacid changes with respect to the mut3 GFP: S30R, Y39N, N105T, Y145F, I171V, and A206V [1].
If there is a need for faster responses in the system, and in consequence, for faster recoveries, we must increase the protein degradation rate. However, we must consider that a strong increase on the degradation rate will imply a loss of accuracy on the reported measurements, as the range will become narrower. Therefore, the addition of the ASV tag becomes ideal as it is supposed to be weak [2].
This ASV tag addition can be seen as an improvement of an existing part as it will make a difference for any kind of system that needs faster continuous measurements. This part is Biobrick compatible.
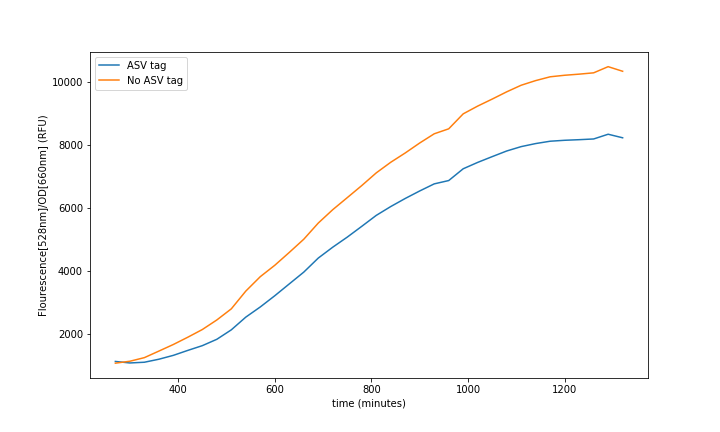
The characterization of this protein was done combining a constitutive promoter (BBa_J23100), a RBS32 (BBa_B0032) and a double terminator (BBa_B0015) followed by the sfGFP with the ASV degradation tag (This reporter sequence BBa_K3484006) . A control with the original sequence (BBa_I746916), with the same promoter, double terminator and RBS, was done to see how the tag affected the final fluorescence.
[1] Pédelacq, J., Cabantous, S., Tran, T. et al. Engineering and characterization of a superfolder green fluorescent protein. Nat Biotechnol 24, 79–88 (2006).
[2] Andersen, J.,Sternberg, C., Poulsen, L. K., Bjørn, S. P., Givskov, M. New Unstable Variants of Green Fluorescent Protein for Studies of Transient Gene Expression in Bacteria. Applied and environmental microbiology, June 1998, p. 2240–2246
Model Characterization
In order to quantify how the ASV tag affected the degradation rate of the sfGFP, a simple mathematical model was developed. The following model is exactly the same for the two scenarios, and consists of a constitutive expression minus a degradation rate (Equation.1). As both systems are constitutively regulated by the same promoter, the PJ23100 (BBa_J23100), the production rate will be the same in the two systems. In addition, the equations representing the steady-state (dy/dx=0) of the system were calculated (Equation.2).
With the Steady-State equations we are able to look for the relationship between the two degradation rates (δ) as the production rate is the same(Equation.3).
In the relation between the degradation rates (Equation 3.) we can clearly see that the ASV tag has an impact on the degradation rate. Mathematically it has been proved that the degradation rate is increased, approximately, 25% when the ASV tag is added on the sfGFP gene sequence. All in all, it has been demonstrated that the addition of the ASV tag increases the degradation rate. Therefore, the introduction of an ASV tag is a good approach to reduce the recovery system time without losing effectiveness of measurement, which is interesting for any kind of biosensor or feedback loop application.
Characterization experiments
For the characterization a Plate-Reader analysis was made. All the information on the experimental conditions and parameters used are described on the table below (Table 1).
| Table 1. Plate-Reader Parameters for the characterization of the effects of ASV tag in sfGFP | |||
| Parameters | Value | ||
| Plate-Reader model | Synergy HTX | ||
| Plate type | Thermo Fischer 96-well microplates black-walled clear bottom | ||
| Cell medium | LB | ||
| Time | 24 hours | ||
| Shake | Linear: Continuous, Frequency: 567 rpm (3mm) | ||
| Temperature | 37C | ||
| Gain | 50 | ||
| Optical Density (OD) measurement (absorbance) | 660nm | ||
| GFP excitation wavelength | 485nm | ||
| GFP emission wavelength | 528nm | ||
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 25
Illegal AgeI site found at 148 - 1000COMPATIBLE WITH RFC[1000]
| None |